Introduction to Lewis and Clark Clusters
This page describes several ways to manage resources and also submit and monitor jobs in a high performance computing (HPC) environment. This is the topic for our "Intro to Linux & Lewis/Clark Cluster" training. As basic Linux is a requirement of using the cluster, the first part of the training will focus on basic Linux commands and the second half of the training focuses on applying these to use on the clusters.
- "Intro to Linux & Lewis/Clark Cluster 2021 PDF"
- "Intro to Linux & Lewis/Clark Cluster 2021 Video"
- Supplemental material on Lewis & Clark Cluster in directory: /group/training/
To be notified about our coming training, please subscribe to our Announcement List.
Older Trainings: - "Intro to Basic Linux 2020 PDF" - "Intro to Basic Linux 2020 Video" - "Intro to Lewis and Clark Clusters 2020 PDF" - "Intro to Lewis and Clark Clusters 2020 Video"
Recommended External Trainings: - Software Carpentry workshop, "The Unix Shell"
RSS Clusters
Clark cluster
- A small-scale cluster for teaching and learning
- No need for registration and available to all MU members by username and PawPrint
- No cost for MU members
Lewis cluster
- A large-scale cluster for requesting high amount of resources
- Great for parallel programming
- GPU resources
- No cost for MU members for general usage
- Investment option is available to receive more resource (more fairshare)
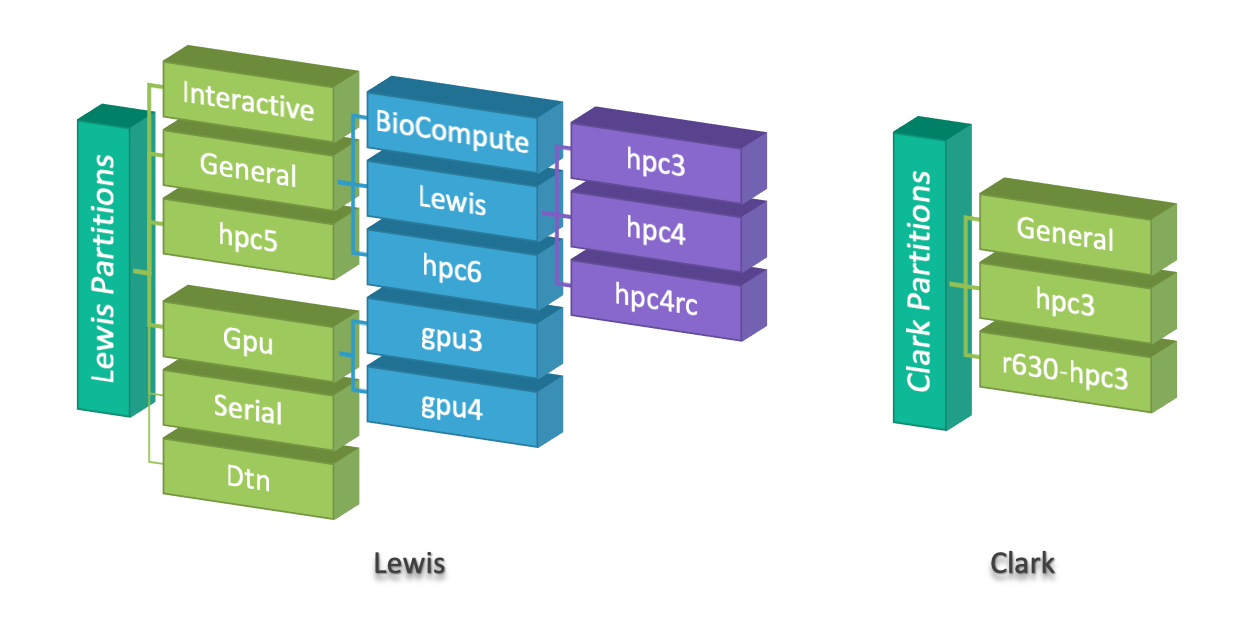
Please see Partition Policy for more information.
Clark and Lewis clusters are accessible by SSH:
- Clark:
ssh <username>@clark.rnet.missouri.edu - Lewis:
ssh <username>@lewis.rnet.missouri.edu
See Getting Started to learn more about accessing RSS clusters.
Slurm
Slurm is an open source and highly scalable cluster management and job scheduling system for large and small Linux clusters. As a cluster workload manager, Slurm has three key functions.
- First, it allocates access to resources (compute nodes) to users for some duration of time so they can perform work
- Second, it provides a framework for starting, executing, and monitoring work (normally a parallel job) on the set of allocated nodes
- Finally, it arbitrates contention for resources by managing a queue of pending work (from Slurm overview)
Moreover,
- All RSS clusters use Slurm
- All Slurm commands start with letter “s"
- Resource allocation depends on your fairshare i.e. priority in the queue
Login Node
All users connect to Clark and Lewis clusters through the login nodes.
[user@lewis4-r630-login-node675 ~]$
[user@clark-r630-login-node907 ~]$
All jobs must be run using Slurm submitting tools to prevent running on the Lewis login node. Jobs that are found running on the login node will be immediately terminated followed up with a notification email to the user.
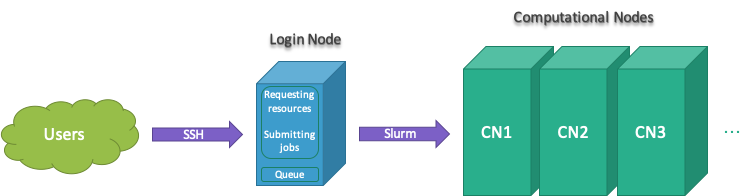
Cluster Information
Slurm is a resource management system and has many tools to find available resources in the cluster. Following are set of Slurm commands for that purpose:
sinfo -s # summary of cluster resources (-s --summarize)
sinfo -p <partition-name> -o %n,%C,%m,%z # compute info of nodes in a partition (-o --format)
sinfo -p Gpu -o %n,%C,%m,%G # GPUs information in Gpu partition (-p --partition)
sjstat -c # show computing resources per node
scontrol show partition <partition-name> # partition information
scontrol show node <node-name> # node information
sacctmgr show qos format=name,maxwall,maxsubmit # show quality of services
./ncpu.py # show number of available CPUs and GPUs per node
For example the following shows output for sinfo -s and sjstat -c commands:
[user@clark-r630-login-node907 ~]$ sinfo -s
PARTITION AVAIL TIMELIMIT NODES(A/I/O/T) NODELIST
r630-hpc3 up 2-00:00:00 0/4/0/4 clark-r630-hpc3-node[908-911]
hpc3 up 2-00:00:00 0/4/0/4 clark-r630-hpc3-node[908-911]
General* up 2:00:00 0/4/0/4 clark-r630-hpc3-node[908-911]
[user@clark-r630-login-node907 ~]$ sjstat -c
Scheduling pool data:
-------------------------------------------------------------
Pool Memory Cpus Total Usable Free Other Traits
-------------------------------------------------------------
r630-hpc3 122534Mb 24 4 4 4
hpc3 122534Mb 24 4 4 4
General* 122534Mb 24 4 4 4
For instance above sinfo -s shows partition hpc3 has 4 idle nodes (free) and users can run jobs up 2 days. And, sjstat -c shows that partition hpc3 has 4 nodes with 24 cpus and 122GB of memory on each node. In general, CPUS/NODES(A/I/O/T) count of CPUs/nodes in the form "available/idle/other/total” and S:C:T counts number of “sockets, cores, threads”.
ncpu.py is a program developed by RSS team that shows CPU and GPU availability at each partition. ncpu.py can be found at /group/training/hpc-intro/alias/.
Users Information
Users can use Slurm to find more information about their accounts, fairshare and quality of services (QOS) and several Unix commands to find their storage quotas.
sshare -U # show your fairshare and accounts (-U --Users)
sacctmgr show assoc user=$USER format=acc,user,share,qos,maxj # your QOS
groups # show your groups
df -h /home/$USER # home storage quota (-h --human-readable)
lfs quota -hg $USER /storage/hpc # data/scratch storage quota (-g user/group)
lfs quota -hg <group-name> /storage/hpc # data/scratch group storage quota
./userq.py # show user’s fairshare, accounts, groups and QOS
Note that:
- Resource allocation depends on your fairshare. If your fairshare is 0.000000 you have used the cluster more than your fair share and will be de-prioritized by the queuing software
- Users have 5GB at their home directory
/home/$USERand 100GB at/storage/hpc/data/$USER - Do not use home directory for running jobs, storing data or virtual environments
- Clark users have 100G on their home storage. The above methods does not apply for Clark
- The RSS team reserves the right to delete anything in
/scratchand/local/scratchat any time for any reason - There are no backups of any storage. The RSS team is not responsible for data integrity and data loss. You are responsible for your own data and data backup
- Please review Storage Policy for more information
userq.py is a program developed by RSS team that shows user’s fairshare, accounts, groups and QOS. userq.py can be found at /group/training/hpc-intro/alias/.
Job Submission
All jobs must be run using srun or sbatch to prevent running on the Lewis login node. In general, users can request resources and run tasks interactively or create a batch file and submit their jobs. The following graphs shows job submission workflow:
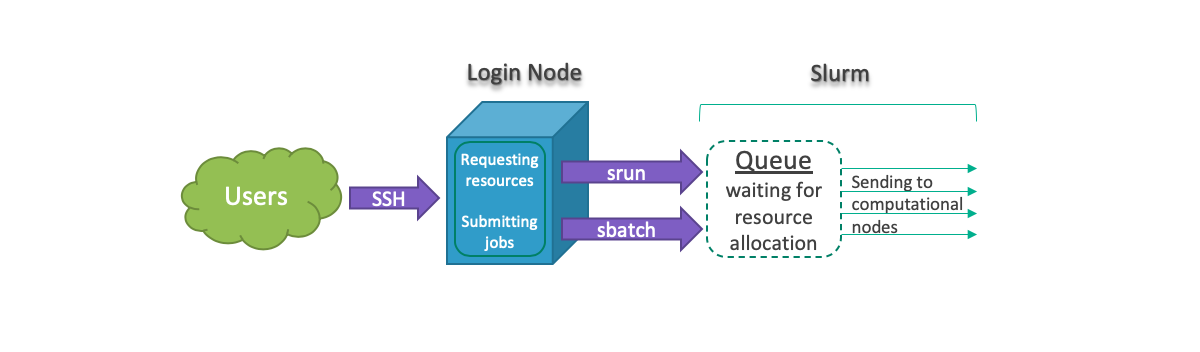
For running jobs interactively, we can use srun to request required resources through Slurm and run jobs interactively. For instance:
srun <slurm-options> <software-name/path>
srun --pty /bin/bash # requesting a pseudo terminal of bash shell to run jobs interactively
srun -p hpc3 --pty /bin/bash # requesting a pseudo terminal of bash shell on hpc3 partition
srun -p Interactive --qos interactive --pty /bin/bash # requesting a p.t. of bash shell in Interactive Node on Lewis (-p --partition)
srun -p <partition-name> -n 4 --mem 16G --pty /bin/bash # req. 4 tasks and 16G memory (-n --ntasks)
srun -p Gpu --gres gpu:1 -N 1 --ntasks-per-node 8 --pty /bin/bash # req. 1 GPU and 1 node for running 8 tasks on Gpu partition (-N --nodes)
For submitting jobs, we can create a batch file, which is a shell script (#!/bin/bash) including Slurm options (#SBATCH) and computational tasks, and use:
sbatch <batch-file>
After job completion we will receive outputs i.e. slurm-jobid.out.
Slurm has several options that help users manage their jobs requirement, such that:
-p --partition <partition-name> --pty <software-name/path>
--mem <memory> --gres <general-resources>
-n --ntasks <number of tasks> -t --time <days-hours:minutes>
-N --nodes <number-of-nodes> -A --account <account>
-c --cpus-per-task <number-of-cpus> -L --licenses <license>
-w --nodelist <list-of-node-names> -J --job-name <jobname>
Also, Slurm has several environmental variables that contain details such as job id, job name, host name and more. For instance:
$SLURM_JOB_ID
$SLURM_JOB_NAME
$SLURM_JOB_NODELIST
$SLURM_CPUS_ON_NODE
$SLURM_SUBMIT_HOST
$SLURM_SUBMIT_DIR
For example let's consider the following Python code, called test.py:
#!/usr/bin/python3
import os
os.system("""
echo hostname: $(hostname)
echo number of processors: $(nproc)
echo data: $(date)
echo job id: $SLURM_JOB_ID
echo submit dir: $SLURM_SUBMIT_DIR
""")
print("Hello world”)
To run the above code, we can use srun to run test.py interactively such that:
srun -p Interactive --qos interactive -n 4 --mem 8G --pty bash # in Lewis
# srun -p hpc3 -n 4 --mem 8G --pty bash # in Clark
python3 test.py
Or create a batch file, called jobpy.sh, such that:
#!/bin/bash
#SBATCH -p hpc3
#SBATCH -n 4
#SBATCH --mem 8G
python3 test.py
And use sbatch to submit the batch file:
sbatch jobpy.sh
Job Arrays
If you have a plan to submit large number of jobs at the same time in parallel, Slurm job array can be very helpful. Note that in most of the parallel programming methods we submit a single script to run but in a way that each iteration of running the same code return different results.
Job arrays use the following environment variables:
SLURM_ARRAY_JOB_IDkeeps the first job ID of the arraySLURM_ARRAY_TASK_IDkeeps the job array index valueSLURM_ARRAY_TASK_COUNTkeeps the number of tasks in the job arraySLURM_ARRAY_TASK_MAXkeeps the highest job array index valueSLURM_ARRAY_TASK_MINkeeps the lowest job array index value
Example 1 - Python
The following shows a simple example of using Slurm job array and Python to submit jobs in parallel. Each of these jobs get a seperate memory and CPU allocation to run tasks in parallel. Let's create the following Python script called array-test.py:
#!/usr/bin/python3
import os
## Computational function
def comp_func(i):
host = os.popen("hostname").read()[:-1] # find host's name
ps = os.popen("cat /proc/self/stat | awk '{print $39}'").read()[:-1] # find cpu's id
return "Task ID: %s, Hostname: %s, CPU ID: %s" % (i, host, ps)
## Iterate to run the computational function
task_id = [int(os.getenv('SLURM_ARRAY_TASK_ID'))]
for t in task_id:
print(comp_func(t))
The following is a batch file called array-job.sh:
#!/usr/bin/bash
#SBATCH --partition hpc3
#SBATCH --job-name array
#SBATCH --array 1-9
#SBATCH --output output-%A_%a.out
python3 ./array-test.py
You can submit the batch file by:
sbatch array-job.sh
The output is:
[user@clark-r630-login-node907 hpc-intro]$ cat output-*
Task ID: 1, Hostname: clark-r630-hpc3-node908, CPU ID: 0
Task ID: 2, Hostname: clark-r630-hpc3-node908, CPU ID: 2
Task ID: 3, Hostname: clark-r630-hpc3-node908, CPU ID: 4
Task ID: 4, Hostname: clark-r630-hpc3-node908, CPU ID: 6
Task ID: 5, Hostname: clark-r630-hpc3-node908, CPU ID: 8
Task ID: 6, Hostname: clark-r630-hpc3-node908, CPU ID: 10
Task ID: 7, Hostname: clark-r630-hpc3-node908, CPU ID: 12
Task ID: 8, Hostname: clark-r630-hpc3-node908, CPU ID: 14
Task ID: 9, Hostname: clark-r630-hpc3-node908, CPU ID: 16
Example 2 - R
Let's use job arrays to rerun the following R script (array-test.R) simultaneously with different seeds (starting point for random numbers):
#!/usr/bin/env R
system("echo Date: $(date)", intern = TRUE)
args = commandArgs(trailingOnly = TRUE)
myseed = as.numeric(args[1])
set.seed(myseed)
print(runif(3))
The following is a batch file called array-job-r.sh:
#!/usr/bin/bash
#SBATCH --partition Lewis
#SBATCH --job-name r-seed
#SBATCH --array 1-12%4 # %4 will limit the number of simultaneously running tasks from this job array to 4
#SBATCH --output Rout-%A_%a.out
module load r
Rscript array-test.R ${SLURM_ARRAY_TASK_ID}
You can submit the batch file by:
sbatch array-job-r.sh
The output is:
[user@lewis4-r630-login-node675 test-array]$ cat Rout-*
[1] "Date: Tue Feb 16 18:21:15 CST 2021"
[1] 0.5074782 0.3067685 0.4269077
[1] "Date: Tue Feb 16 18:21:15 CST 2021"
[1] 0.2772497942 0.0005183129 0.5106083730
[1] "Date: Tue Feb 16 18:21:15 CST 2021"
[1] 0.06936092 0.81777520 0.94262173
[1] "Date: Tue Feb 16 18:21:15 CST 2021"
[1] 0.7103224 0.2461373 0.3896344
Monitoring Jobs
The following Slurm commands can be used to monitor jobs:
sacct -X # show your jobs in the last 24 hours (-X --allocations)
sacct -X -S <yyyy-mm-dd> # show your jobs since a date (-S --starttime)
sacct -X -S <yyyy-mm-dd> -E <yyyy-mm-dd> -s <R/PD/F/CA/CG/CD> # show running/pending/failed/cancelled/completing/completed jobs in a period of time (-s --state)
sacct -j <jobid> # show more details on selected jobs (-j --jobs)
squeue -u <username> # show a user jobs (R/PD/CD) in the queue (-u --user)
squeue -u <username> --start # show estimation time to start pending jobs
scancel <jobid> # cancel jobs
./jobstat.py <day/week/month/year> # show info about running, pending and completed jobs of a user within a time period (default is week)
jobstat.py is a program developed by RSS team that shows information about running, pending and completed jobs of a user within a time period (default is one week). jobstat.py can be found at /group/training/hpc-intro/alias/.
Monitor CPU and Memory
Completed jobs
We can use the following options to find efficiency of the completed jobs:
sacct -j <jobid> -o User,Acc,AllocCPUS,Elaps,CPUTime,TotalCPU,AveDiskRead,AveDiskWrite,ReqMem,MaxRSS # info about CPU and virtual memory for compeleted jobs (-j --jobs)
seff <jobid> # show job CPU and memory efficiency
For more information on what fields to include see man sacct.
Example output:
[user@lewis4-r630-login-node675 ~]$ sacct -j 10785018 -o User,Acc,AllocCPUS,Elaps,CPUTime,TotalCPU,AveDiskRead,AveDiskWrite,ReqMem,MaxRSS
User Account AllocCPUS Elapsed CPUTime TotalCPU AveDiskRead AveDiskWrite ReqMem MaxRSS
--------- ---------- ---------- ---------- ---------- ---------- -------------- -------------- ---------- ----------
user general 16 00:48:39 12:58:24 01:49.774 66.58M 44.75M 64Gn 216K
[user@lewis4-r630-login-node675 ~]$ seff 10785018
Job ID: 10785018
Cluster: lewis4
User/Group: user/rcss
State: COMPLETED (exit code 0)
Nodes: 1
Cores per node: 16
CPU Utilized: 00:01:50
CPU Efficiency: 0.24% of 12:58:24 core-walltime
Memory Utilized: 3.38 MB (estimated maximum)
Memory Efficiency: 0.01% of 64.00 GB (64.00 GB/node)
Running jobs
Slurm also, provides sstat tool to monitor *running jobs efficiency. In order for this to work, jobs using SBATCH must utilize the srun command within the job file.
sstat <jobid> -o AveCPU,AveDiskRead,AveDiskWrite,MaxRSS # info about CPU and memory for runing jobs (srun only)
Moreover, we can use top command to find how much CPU and memory are using by a running job. To do that, we need to attach to the node that our job is running and use top command by using:
srun --jobid <jobid> --pty /bin/bash
top -u $USER
For Memory usage, the number you are interested in is RES. In the below example, python3 program is using about 5.6Mb memory and 0% of requested CPUs.
PID USER PR NI VIRT RES SHR S %CPU %MEM TIME+ COMMAND
14278 rcss 20 0 124924 5612 2600 S 0.0 0.0 0:00.04 python3
14279 rcss 20 0 124924 5612 2600 S 0.0 0.0 0:00.03 python3
Modules
In order to use a software, we need to load the corresponding module first. The following commands let us manage modules in our workflow:
module avail # available modules
module show # show modules info
module list # list loaded modules
module load # loaded modules
module unload # unload loaded modules
module purge # unload all loaded modules
Never load modules in the login node. It makes login node slow for all users and many modules don’t work in the login node.
For example to use R interactively, first need to request resources by srun and then use module load R:
srun -p Interactive --qos interactive --mem 4G --pty /bin/bash
# srun -p hpc3 --mem 4G --pty bash # in Clark
module load R
R
If you are looking for using a licensed software (available in cluster) make sure you call the license when requesting resources. For instance to use MATLAB:
srun -p Interactive --qos interactive --mem 4G -L matlab --pty /bin/bash
module load matlab
matlab -nodisplay
Review our Software documentation to find more details about running a software in our clusters. Review Environment Modules User Guide for more details.